Hello,
I am trying to use VirtualFlow on the Texas Advanced Computing Center (TACC) but I have run into a job configuration error and a subsequent memory error. I am unsure if the two are related, so I will discuss them both here.
TACC is configured such that you cannot submit a job from a compute node (only a login node), so I am trying to avoid using that functionality. Moreover, on TACC the preferred approach is to submit one job with many nodes, instead of many jobs with one/few nodes. I believe VirtualFlow can still be configured to meet these needs, so I will outline my general approach here.
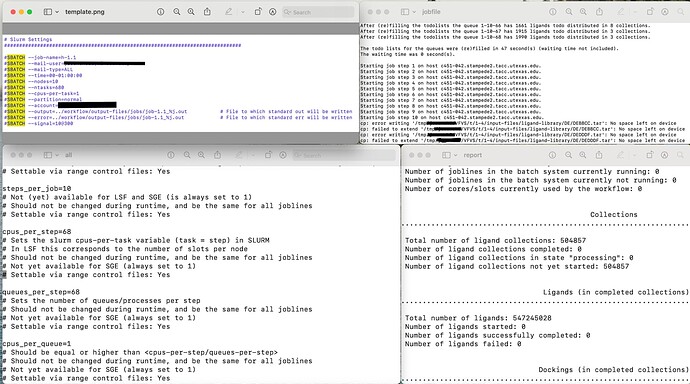
Attached are relevant screenshots of the .ctrl and .slurm files.
These produced the following error when I submitted them using ./vf_start_jobline.sh:
- sbatch: error: Batch job submission failed: Requested node configuration is not available
However, I was able to circumvent this error when I commented out the --cpus-per-task #SBATCH directive. Notably, when I simply delete the directive instead of commenting it out, I also obtain an error and the job does not enter the queue. Why is that?
Also, when I analyze the workflow via ./vf_report.sh (on the job with the --cpus-per-task directive commented out) I notice that none of the ligands were processed (see attached report screenshot).
Of course, I looked at the job file (also attached) to see if I could glean what the issue was. It seems to be related to memory/storage. I cannot think of how to remedy this, though.
Also, from the job file, I noticed that there 10 job steps, but they are all appearing to execute on the same node (c451-042).
i.e.:
Starting job step 1 on host c451-042.stampede2.tacc.utexas.edu
Starting job step 2 on host c451-042.stampede2.tacc.utexas.edu
…
Starting job step 10 on host c451-042.stampede2.tacc.utexas.edu
My intention is to spread out computational and memory loads against the nodes that request. I am not sure if I did something wrong in the .ctrl or the .slurm files, though.
I would really appreciate help if anyone has a solution. I think VirtualFlow is a fantastic tool and I look forward to using it on my research!